Build a Fungus Foraging App with Machine Learning
Follow articleHow do you feel about this article? Help us to provide better content for you.
Thank you! Your feedback has been received.
There was a problem submitting your feedback, please try again later.
What do you think of this article?
Whilst in the height of the 2019 mushroom foraging season I chose to combine my thirst for knowledge about low-level machine learning (ML) with a popular pastime that we enjoy here on Anglesey. Just for the record, I’m not an expert in ML, coding or fungi, and I’m simply inviting readers to follow me back down some rabbit holes that I recently explored.
Firstly, a bit about health and safety:
|
|
Since this is very much an introduction to ML, there won’t be too much terminology and the emphasis will be on having fun rather than going on a mind-boggling deep dive. The system that I stumbled upon is called XGBoost (XGB). One of the XGB demos is for binary classification, and the data was drawn from The Audubon Society Field Guide to North American Mushrooms. Binary means that the app spits out a probability of ‘yes’ or ‘no’ and in this case it tends to give about 95% probability that a common edible mushroom (Agaricus campestris) is actually edible, which is reassuringly inaccurate!
The app asks the user 22 questions about their specimen and collates the data inputted as a series of letters separated by commas. At the end of the questionnaire, this data line is written to a file called ‘fungusFile.data’ for further processing.
XGB can not accept letters as data so they have to be mapped into ‘classic LibSVM format’ which looks like this: ‘3:218’, for each letter. Next, this XGB friendly data is split into two parts for training a model and then subsequently testing that model.
Installing XGB is relatively easy compared to higher-level deep learning systems and runs well on both Linux Ubuntu 16.04 and on a Raspberry Pi. I wrote the deployment app in bash so there should not be any additional software to install. Before getting any deeper into the ML side of things, I highly advise installing XGB, running the app, and having a bit of a play with it. Machine learning can definitely be fun!
Training and testing is carried out by running bash runexp.sh in the terminal and it takes less than one second to process the 8124 lines of fungal data. At the end, bash spits out a set of statistics to represent the accuracy of the training and also attempts to ‘draw’ the decision tree that XGB has devised. If we have a quick look in the directory, ~/xgboost/demo/binary_classification, there should now be a 0002.model file in it ready for deployment with the questionnaire.
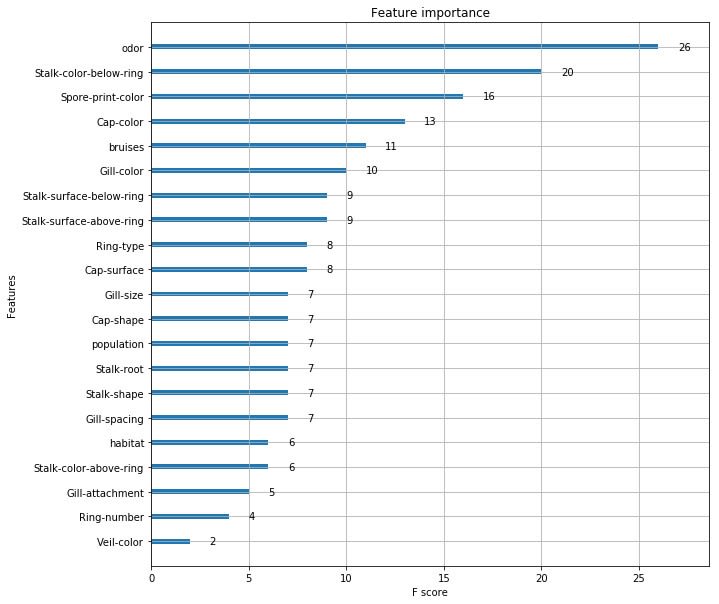
Rather than blindly trusting the ML algorithms, it's always a good idea to try and explore the classification results a bit further and, in this case, look at the way XGB weighted different characteristics of the fungi. I eventually got some rough visualisations working on a Python-based Jupyter Notebook script:
Obviously, this app is not going to win any online competitions such as Kaggle as the various parameters within the software need to be carefully tuned with the help of all the different software tools available. A good place to start is to tweak the maximum depth of the tree and the number of trees used. Depth = 4 and number = 4 seems to work well for this data. Other parameters include the feature importance type, for example, gain, weight, cover, total_gain or total_cover. These can be tuned using tools such as SHAP. Finally, this app could easily be adapted to other questionnaire-based systems such as diagnosing a particular disease or deciding whether to buy a particular stock or share in the market place.
If you're now inspired to explore ML a bit further, check out my next article on ML for ultrasonic audio - Watch this space!